This is part 2 of an article based on a presentation prepared for an Schering-Plough Animal Health roadshow about salmonella, sources of contamination, laboratory testing, the diseases and control.
Table of Contents
Knowing your enemy
- Salmonellae are bacteria that can cause infectious disease
- Other causes of infectious disease include (smallest to largest):
- Viruses
- Chlamydia
- Coccidia
- Bacteria
- Worms
- Mycoplasma
- Mites
- Fungi
Bacteria are very small and can live on or in cells of the body. They are characterised by a solid outer wall to which are attached flagellae. These enable the bacterium to move and smaller ones stick the bacterium to cell surface. They can be seen using a light microscope. Using special stains can make this easier. Salmonellae can be cultured in the laboratory.
Chlamydia and mycoplasmas are specialised types of bacteria. Bacteria are the smallest living infectious organisms.
Viruses are non-living. These need the living cells to replicate, bacteria, the chicken. They effectively hijack the cell so that rather than make normal cell components, viral components are made instead. Often this can result in cell death.
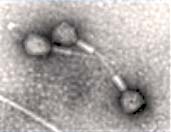
Viruses can infect bacteria. The black and white picture is an electron micrograph of bacteriophages that infect salmonllae. The picture below shows salmonellae growing on the right.
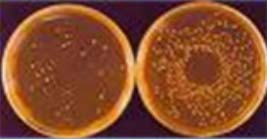
If the cultures are infected with a bacteriophage the salmonellae die, fail to grow so there are less to be seen. These viral bacteriophages are used to further identify the salmonella serovars. This helps track infections and bacterial changes.
Can you identify your enemy
Scientist tell us that salmonellae belong to:
| Science | Meaning |
| The family Enterobacteriacaea | Intestinal bacteria – E.coli, Proteus sp. |
| Gram Negative | Stain red |
| Aerobic or facultatively anaerobic | Grow best with oxygen |
| Asporogenous | Do not produce spores |
| Rod shaped | Not spherical |
| Grow on artificial media | Can be cultured |
So we can identify their presence and separate them from other gut bacteria.
| Heat | 5 – 45c. Roasting, boiling but soft yolks a problem. |
| pH | 4.0 – 9.0 optimum 7.0 |
| UV | Lethal |
| Litter | Survive 7 – 16 months at 20 – 25c |
| Feed | Survive 18 months |
| Chemicals | Most effective is formaldehyde, others help |
S.Senftenberg are more heat resistant than most. Internal temperature of 79C kills St in roasting chickens. 60C for 5 minutes killed 100 million bacteria per gram of ground chicken meat. Prior refrigeration increases susceptibility of Se to temperature.
Ultrasound, irradiation and electrical stimulation also help.
Salmonellae lab test classifications
Into 2 species:
- S.enterica
- This is futher divided into 6 sub species e.g
- Host adapted, non-motile – S.Pullorum, S.Gallinarum
- Non Host adapted, motile 2500+ serovars which includes
- S.enterica subsp. enterica serovar Typhimurium
- This is futher divided into 6 sub species e.g
- More commonly known as Se or St
The import species for us is S.enterica. This species is further subdivided into 6 subspecies. The important one for us being enterica. Thus officially St would be known as S.enterica subsp enterica serovar Typhimurim / Enteritidis / Virchow as appropriate. These are still usually shortened to Se and St.
The importance of being able to differentiate salmonellae using specific antibodies enables us to study how the infection spreads and develops. For instance Se belongs to serological group D and St to Group B.
Salmonella identification
- Salmonella enterica subsp enterica serovar Pullorum (Sp)
- Disease of chicks eradicated from commercial flocks.
- S.Gallinarum (Sg)
- Disease of laying birds eradicated from commercial flocks.
- S.Enteritidis (Se)
- Opportunistic pathogen of fowl, commonest cause of salmonella food poisoning in humans.
This group also includes Pollorum Disease, Fowl Typhoid, S.Dublin (a cattle salmonella) and many others. Using these antibodies enables us to distinguish between the subtle differences on the surface of these bacteria.
In the lab we can culture salmonellae and identify which of the 2500 we are dealing with.
